Research
Single-molecule dynamics in live cells
Biologists have known the existence of biomolecules for ages; yet it is in this only in this century has it become feasible to directly visualize individual biomolecules in a living specimen using modern fluorescence microscopy techniques. One of the lab’s research focus is to apply single-molecule imaging to study real-time dynamics of complicated molecular interactions in cells. We are particularly interested in understanding how protein molecules’ behavior changes during signal transduction, and how individual proteins are made during gene expression.
For more detail please see the following publications from the lab:
Super resolution optical microscopy
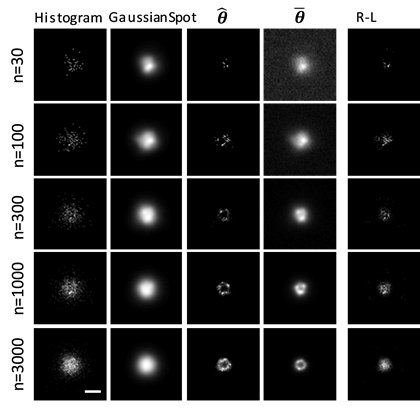
The invention of fluorescence microscopy in the early twenties century has had a tremendous impact on the advancement of biology research, as it allowed the biologist to define cellular or sub-cellular structures in a molecule-specific fashion. Nevertheless, for a long time fluorescence microscopy had suffered from the curse of the so-called “diffraction-limited resolution”, which sets a fundamental limit on what is the smallest object the microscope can see. In the last decade, however, many new forms of microscopy design have emerged with the goal of breaking this resolution limit. Collectively, they are often called “super-resolution optical microscopy”. The lab is actively working on this new filed trying to further improve the imaging quality of these new technologies.
For more detail, check out these recent papers: